library(tidyverse)
library(repurrrsive) # devtools::install_github("jennybc/repurrrsive") # or install.packages("repurrrsive")
For Each Do you are already solving them: copy & paste, for loops, (l/s)apply()
Alternative purrr::map() & friends
library(purrr)
str(sw_people[1:3])
map()¶
map(.x, .f, ...)
for each element of .x do .f
- .x: a vector, a list, a data frame(for each column)
- .f: We'll get to that...
How many starships has each character been in?¶
for each person in sw_people, count the number of starships
map(swpeople, _ )
Strategy
Do it for one element
Turn it into a recipe
Use
map()to do it for all elements
luke <- sw_people[[1]]
# How many starships has luke been in?
length(luke$starships)
leica <- sw_people[[5]]
length(leica)
Turn it into a recipe¶

map(sw_people[1:5], ~ length(.x$starships))
planet_lookup <- map_chr(sw_planets[1:5],"name") %>%
set_names(map_chr(sw_planets[1:5],"url"))
planet_lookup
Find the name of each characters home world¶
find the body mass index(BMI) of all characters
bmi = (mass in kg)/(height in m)^2)
luke$homeworld
planet_lookup[luke$homeworld]
map(sw_people[1:5], ~planet_lookup[.x$homeworld])
ROADmap()¶
map_lgl(.x, .f, ...)
Other types of output
Other ways of specifying .f
Other iteration functions
map2(.x, .y, .f, ...)
Other types of output
Other ways of specifying .f
Other iteration functions
map() details¶
map() always returns a list
simple output:
map_lgl(): logical vector
map_int(): integer vector
map_dbl(): double vector
map_chr(): character vector
walk: when you want nothing at all, use a function for its side effects
Result: No surprises!
vector same length as .x or an ERROR
# names can be useful
sw_people <- sw_people %>% set_names(map_chr(sw_people, "name"))
Repalce map() with the appropriately typed function¶
# How many starships has each character been in?
map(sw_people[1:5], ~length(.x[["starships"]]))
map_int(sw_people[1:5], ~ length(.x[["starships"]]))
# what is color is each character's hair?
map(sw_people[1:5], ~ .x[["hair_color"]])
map_chr(sw_people[1:5], ~ .x[["hair_color"]])
# Is the character male?
map(sw_people[1:5], ~ .x[["gender"]] == "male")
# How heavy is each character?
map(sw_people[1:5], ~ .x[["mass"]])
map_lgl(sw_people[1:5], ~ .x[["gender"]] == "male")
# How heavy is each character?
map_dbl(sw_people, ~ .x[["mass"]])
Doesn't work... because we get a string back¶
map(sw_people[1:5], ~ .x[["mass"]])
# A little risky
map_dbl(sw_people[1:5], ~ as.numeric(.x[["mass"]]))
# Probably want something like:
map_chr(sw_people[1:5], ~ .x[["mass"]]) %>%
readr::parse_number(na = "unknown")
(1) .f can be a string or integer¶
for each element, sxtract the named/numbered element
map(.x, .f = "some_name")
equivalent to
map(.x, ~ .x[["some_name"]]
(2) .f can be a function¶
map(.x, .f = some_function, ...)
equivalent to
map(.x, ~ some_function(.x, ...))
char_starships <- map(sw_people[1:5], "starships")
map_int(char_starships, length)
# In one go
map(sw_people[1:5], "starships") %>% map_int(length)
# also equivalent to
map_int(sw_people[1:5], ~ length(.x[["starships"]]))
From earlier...¶
create planet_lookup:
planet_lookup <- map_chr(sw_planets[1:5], "name") %>%
set_names(map_chr(sw_planets[1:5], "url"))
planet_lookup

What about sapply() & lapply()?¶
What type of object does sapply() return? It depends.
Motivation for purrr:
- consistent return type,
- useful shortcuts,
- consistent syntax for more complicated iteration
Star Wars challenges¶
# Which film (see sw_films) has the most characters?
map(sw_films[1:5], "characters") %>%
map_int(length) %>%
set_names(map_chr(sw_films[1:5], "title")) %>%
sort()
# Which species has the most possible eye colors?
sw_species[[1]]$eye_colors
map_chr(sw_species[1:5], "eye_colors") %>%
strsplit(", ") %>%
map_int(length)
# this is lazy, what about n/a and unkown?
to each element of .x apply .f¶
walk(.x, .f )
Expect nothing in return
You actually get .x invisibly back, good for piping

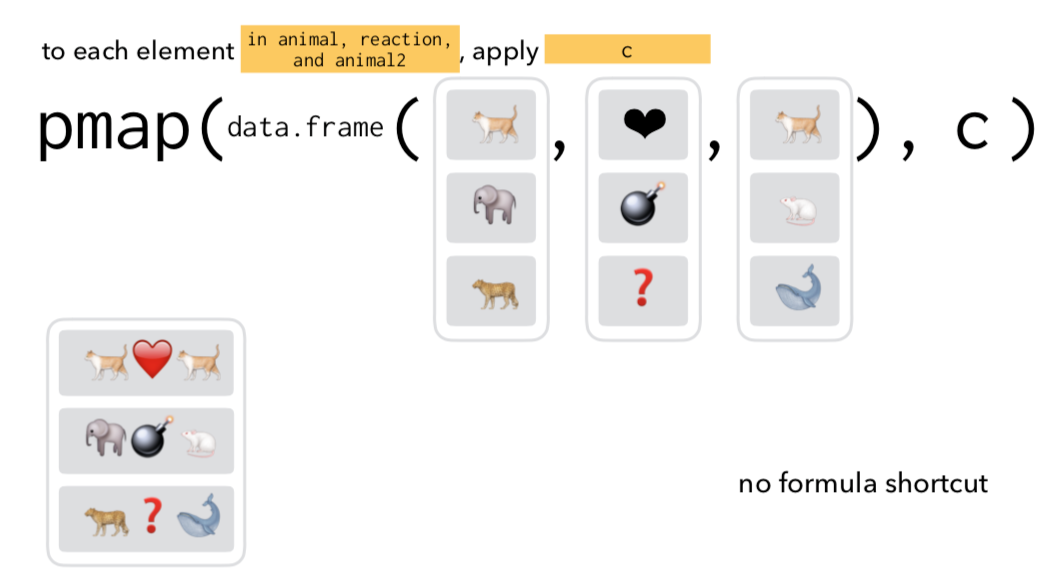
to each element of .x and corresponding element of .y apply .f
map2(.x, .y, .f)

Discuss¶
1.
For each function, which two arguments might be useful to iterate over?
download.file()
rnorm()
lm()
predict.lm()
write.csv()
2.
For which functions above should we use walk2() or a typed version of map2()?
download.file() for each url download to destfile walk2(), map2_int()
rnorm() for each n generate a Normal sample with mean mean (or sd) (See purrr::rerun() for repeating a function many times)
lm() for each data fit a model(formula)
predict.lm() for each model(object), generate predictions at data(newdata)
readr::write_csv() for each data frame(x) save to path. Similar for ggplot::ggsave() for each plot save to filename walk2()
What does this code do?¶
jan_sales <- read_csv("jan.csv")
jan_sales <- mutate(jan_sales, month = "jan")
feb_sales <- read_csv("feb.csv")
feb_sales <- mutate(feb_sales, month = "feb")
mar_sales <- read_csv("mar.csv")
mar_sales <- mutate(mar_sales, month = "mar")
sales <- bind_rows(jan_sales, feb_sales, mar_sales)
Reduce duplication (and mistakes) with purrr¶
months <- c("jan", "feb", "mar")
files <- paste0(months, ".csv")
sales_list <- map(files, read_csv)
Now... For each element(do) add a month column
Use the same strategy!
Do it for one¶
Solve the problem for one element
mutate(sales_list[[1]], month = months[[1]])
mutate(sales_list[[2]], month = months[[2]])
Iterating over two objects!
Do it for all!¶
months <- c("jan", "feb", "mar")
files <- paste0(months, ".csv")
sales_list <- map(files, read_csv)
sales_list_months <- map2(.x = sales_list,
.y = months,
.f = ~mutate(.x, month = .y)
bind_rows(sales_list_months)
library(repurrrsive)
gap_split_small <- gap_split[1:10]
countries <- names(gap_split_small)
For each country create a ggplot of life expectancy through time with a title¶
ggplot(gap_split[[1]], aes(year, lifeExp)) +
geom_line() +
labs(title = countries[[1]])
# For all countres
plots <- map2(gap_split_small, countries,
~ ggplot(.x, aes(year, lifeExp)) +
geom_line() +
labs(title = .y))
plots[[1]]
# Display all plot
walk(plots[1:3], print)
purrr and list columns¶
purrr and list columns
Data should be in a data frame as soon as it makes sense!
Data frame: cases in rows, variable in columns
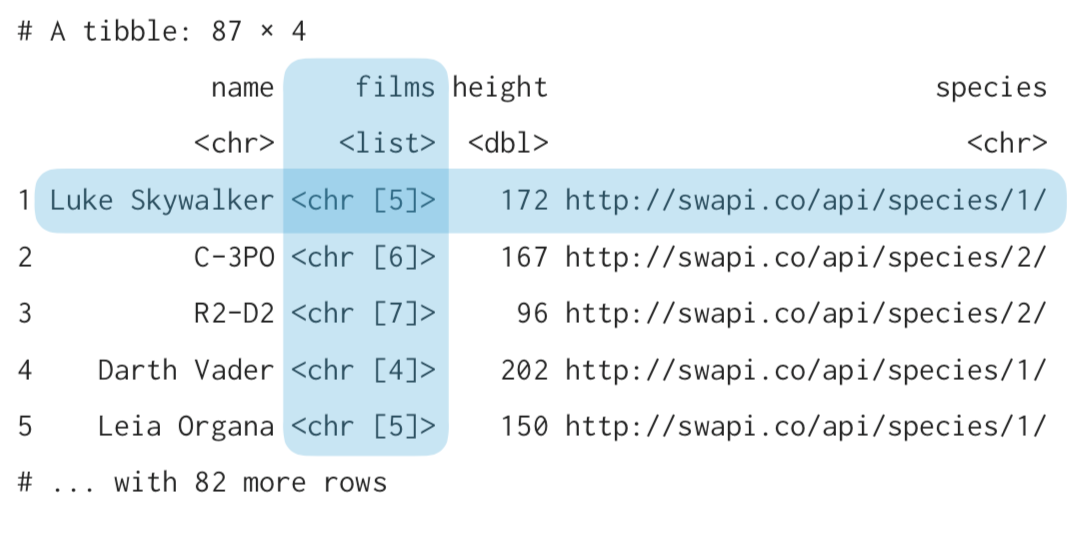

purrr can help lists into tibbles¶
library(tidyverse)
people_tbl <- tibble(
name = sw_people %>% map_chr("name"),
films = sw_people %>% map("films"), #will result in list column
height = sw_people %>% map_chr("height") %>%
readr::parse_number(na = "unknown"), #needs some parsing
species = sw_people %>% map_chr("species", .null = NA_character_)
) #isn't in every element
head(people_tbl,3)
combine purrr with dplyr to work with list columns¶
# A useful lookup table -----------------------------------------------
film_number_lookup <- map_chr(sw_films, "url") %>%
map(~ stringr::str_split_fixed(.x, "/", 7)[, 6]) %>%
as.numeric() %>%
set_names(map_chr(sw_films, "url"))
people_tbl <- tibble(
name = sw_people %>% map_chr("name"),
films = sw_people %>% map("films"),
height = sw_people %>% map_chr("height") %>%
readr::parse_number(na = "unknown"),
species = sw_people %>% map_chr("species", .null = NA_character_)
)
# Turning parts of our list to a tibble ---------------------------------
# people_tbl$films
# Use map with mutate to manipulate list columns
people_tbl <- people_tbl %>%
mutate(
film_numbers = map(films,
~ film_number_lookup[.x]),
n_films = map_int(films, length)
)
people_tbl[1:5,] %>% select(name, film_numbers, n_films)
Create a new character column that collapses the film numbers into a single string, e.g. for Luke:"6,3,2,1,7"
?paste
people_tbl <- people_tbl %>%
mutate(film_squashed = map_chr(film_numbers, paste, collapse = ","))
people_tbl[1:5,] %>% select(name, n_films, film_squashed)
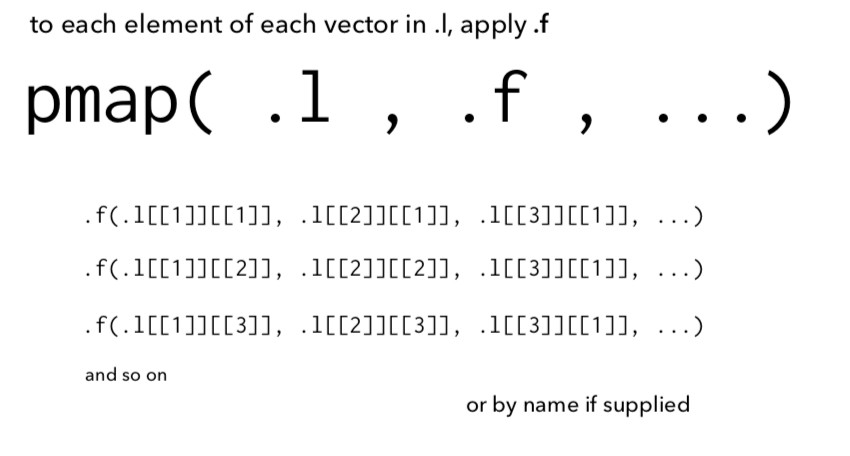



Lists and functions¶
Key objects in purrr
purrr provides a pile of functions to make working with them easier
Functions: safely(), possibly(), partial()
Lists: transpose(),accumulate(),reduce(),every(),order_by()
Wrap up¶
purrr provides:
- functions that write for loops for you
- with consistent syntax, and
- convenient shortcuts for specifying functions to iterate
Choosing the right function depends on:
- type of iteration
- type of output
